🟥 Self-Supervised Divergent Embeddings (SSDE) for Divergence-Driven Stimulus Selection
Introduction
The Red Team track of the Re-Align Challenge 2026 asks participants to submit a set of 1000 images intended to maximize representational divergence across a fixed suite of vision models, with divergence evaluated on a hidden split using Centered Kernel Alignment (CKA) between model embeddings. [@realign_workshop_2026; @realign_leaderboard_space_2026; @kornblith2019cka] CKA is closely related to the Hilbert–Schmidt Independence Criterion (HSIC), providing a robust similarity measure that is invariant to isotropic scaling and commonly used for comparing representational geometry. [@gretton2005hsic; @kornblith2019cka] Our goal is therefore to select stimuli that systematically expose differences in representational geometry across models, rather than simply selecting images that are difficult in a behavioral sense.
We frame stimulus selection as set optimization over a candidate pool drawn from the datasets allowed by the challenge (ImageNet validation and ObjectNet). [@deng2009imagenet; @barbu2019objectnet; @objectnet_website] The approach has two stages:
- Baseline selector (proxy divergence + diversity): compute per-image divergence signatures from multi-model representations, score images using a multi-directional disagreement proxy (including a log-det term), then select a set of 1000 images with an explicit diversity criterion.
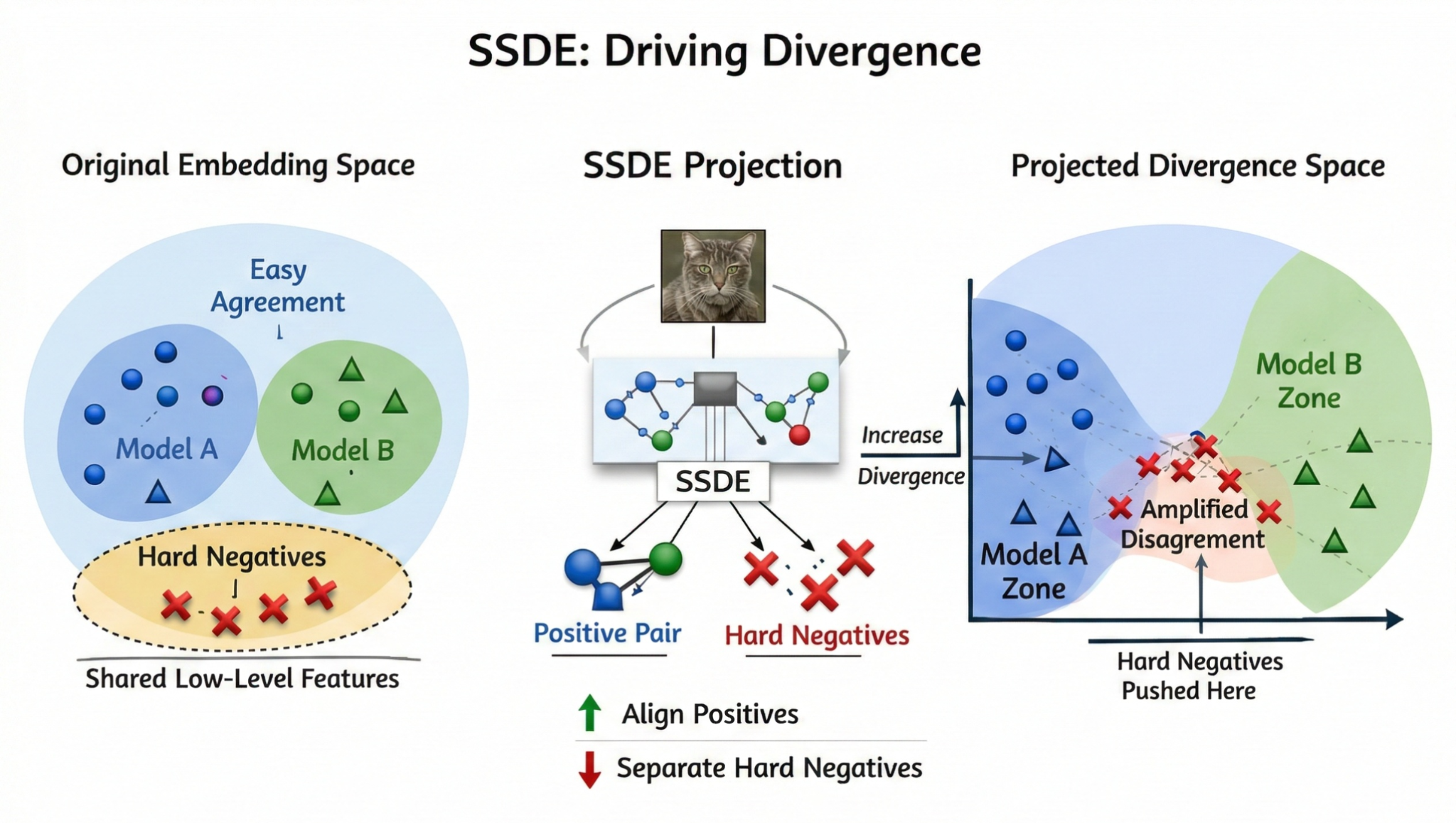
- Enhanced selector (SSDE + diversity): learn a Self-Supervised Divergence Embedding (SSDE) from the signatures using contrastive learning with hard negative mining, then perform diversity-aware selection in the learned embedding space. [@chen2020simclr; @robinson2021hard_negatives; @kalantidis2020hard_negative_mixing]
All experiments were run on a multi-node GPU cluster for representation extraction, with CPU- and GPU-based jobs for aggregation, SSDE training, and subset selection.
Methods
Objective and notation
Let the candidate stimulus pool (of images) be (\mathcal{D}={x_i}{i=1}^N), where each (x_i) is an image from either ImageNet-val or ObjectNet. Let ({f_m}{m=1}^M) denote the suite of (M) fixed vision models used by the challenge. For each model (m), we extract a feature vector at a designated layer: [ h_m(x_i)\in\mathbb{R}^{d_m}. ] The challenge evaluation computes a representational similarity (CKA) over sets of embeddings across images. [@kornblith2019cka] Because we do not have access to the hidden evaluation set, we build per-image proxies designed to predict which stimuli will decrease cross-model similarity when aggregated.
Stage A — Dimensionality harmonization via random projections
Models produce features of different dimensions (d_m). To standardize computations and reduce I/O and memory, we map each (h_m(x_i)) into a shared dimension (d) using a fixed random projection: [ e_m(x_i) = P_m,\tilde{h}_m(x_i)\in\mathbb{R}^{d}, ] where:
- (\tilde{h}_m(x_i)) is a standardized version of (h_m(x_i)) (e.g., centering and (\ell_2)-normalization),
- (P_m\in\mathbb{R}^{d\times d_m}) is a random matrix sampled once per model and held fixed.
Random projections approximately preserve pairwise distances under mild conditions (Johnson–Lindenstrauss), enabling substantial dimensionality reduction while retaining geometric structure. [@johnson1984jl; @achlioptas2003random_projections]
In our runs we used (d=256) for projected model features (reported in Table 1).
Stage B — Divergence signatures and multi-directional disagreement scores
For each image (x_i), we stack the projected features across models into a matrix:
[
E_i =
\begin{bmatrix}
e_1(x_i)^\top
\vdots
e_M(x_i)^\top
\end{bmatrix}
\in\mathbb{R}^{M\times d}.
]
B.1 Pairwise disagreement (“hardness” proxy)
A simple proxy for cross-model disagreement is the mean pairwise cosine distance: [ u_i ===
\frac{2}{M(M-1)} \sum_{1\le a < b \le M} \left( 1-\frac{\langle e_a(x_i), e_b(x_i)\rangle}{|e_a(x_i)|_2,|e_b(x_i)|_2} \right). ] This increases when different models place the same image in dissimilar directions in feature space, which we expect to reduce representational alignment when many such images are aggregated. This choice is aligned with the representational-similarity viewpoint that comparisons should reflect geometry, not raw coordinates. [@kriegeskorte2008rsa; @sucholutsky2023getting_aligned]
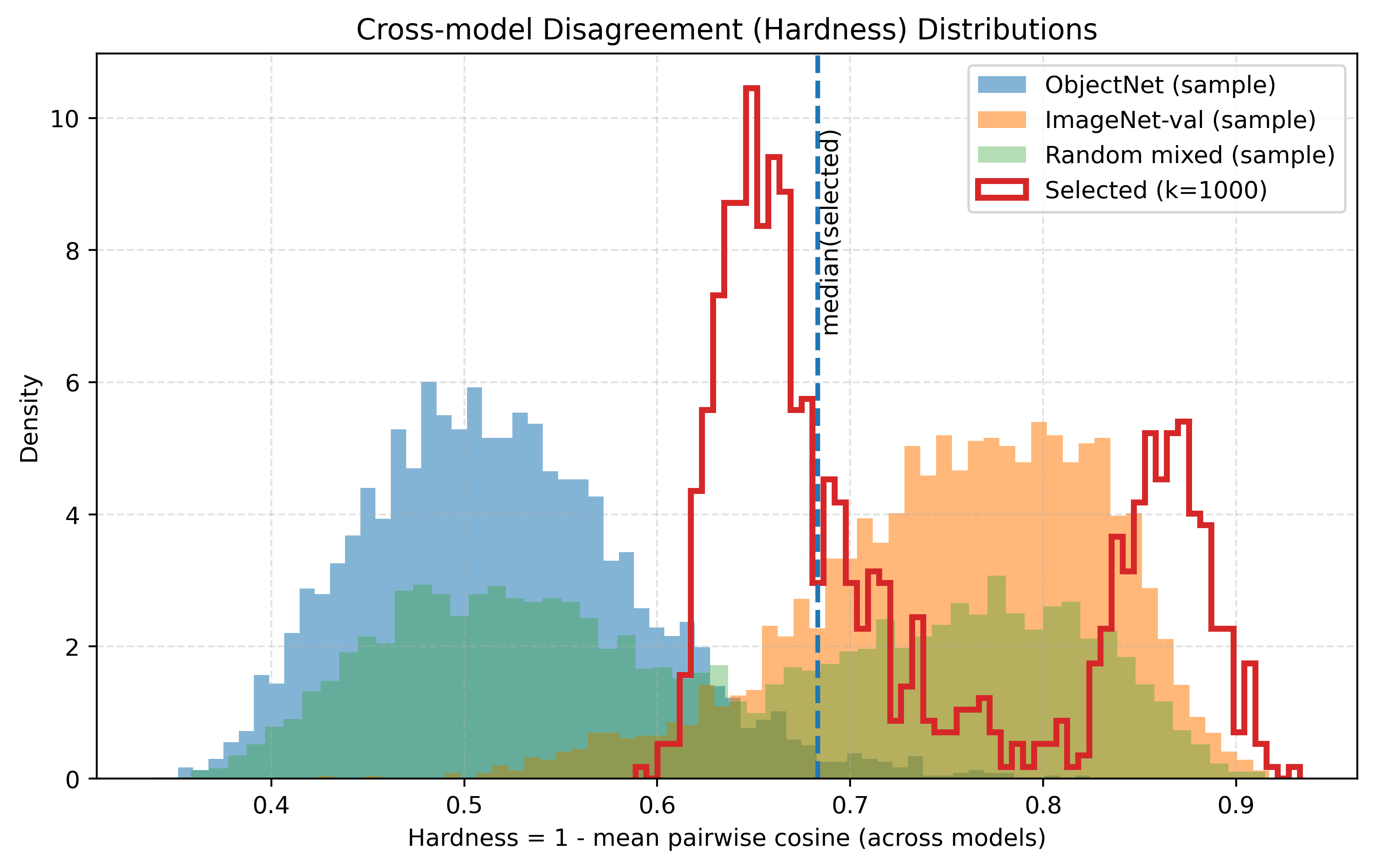
The diagnostic plot in Figure 2 visualizes this quantity as “hardness”: [ \mathrm{hardness}(x_i) \equiv u_i. ]
B.2 Multi-directional disagreement via log-det
Pairwise averages can over-select stimuli that drive disagreement in a single direction (e.g., a shared “failure mode”). We therefore include a multi-directional term based on the log-determinant of the model–model Gram matrix: [ G_i = \frac{1}{d}E_i E_i^\top \in \mathbb{R}^{M\times M}, \qquad \ell_i = \log\det\left(G_i + \varepsilon I\right). ] Here (\ell_i) is large when the model feature vectors for (x_i) span a high-volume simplex (i.e., are diverse and not confined to a low-dimensional subspace). Log-det objectives are widely used as diversity/coverage surrogates and appear in submodular sensor-placement and information-gain settings. [@krause2008sensor_placement]
B.3 Divergence signature and scalar score
We define a divergence signature (s_i\in\mathbb{R}^{D}) that concatenates:
- the upper-triangular entries of (G_i) (flattened), and
- auxiliary summary statistics such as (u_i) and (\ell_i).
A scalar divergence score is then computed as: [ d_i = \alpha,u_i + (1-\alpha),\operatorname{clip}(\ell_i;,\ell_{\min},\ell_{\max}), ] with (\alpha\in[0,1]) controlling the contribution of pairwise vs. multi-directional disagreement and (\varepsilon>0) stabilizing the log-det.
Stage C — Diversity-aware subset selection (baseline selector)
Selecting the top 1000 images by (d_i) alone tends to produce near-duplicates. We therefore treat selection as maximizing a score–diversity trade-off.
C.1 MMR-style greedy selection
Given an embedding (v_i) used solely for measuring redundancy (either (s_i) or SSDE embeddings (z_i); see below), we use a greedy Maximal Marginal Relevance (MMR) criterion: [@carbonell1998mmr] [ i^\star = \arg\max_{i\notin S} \left[ d_i —
\lambda\cdot \max_{j\in S}\mathrm{sim}(v_i, v_j) \right], \qquad \mathrm{sim}(a,b)=\frac{a^\top b}{|a|_2|b|_2}. ] (\lambda\ge 0) is the diversity weight. To keep runtime manageable, selection is performed on the top (P) candidates by (d_i) (a “prefilter”).
C.2 Relation to log-det/DPP diversity (context)
As an alternative diversity regularizer, one can use: [ \log\det(K_S+\delta I), ] where (K) is a positive semidefinite similarity kernel on candidate signatures. This is closely related to determinantal point processes (DPPs), a classic probabilistic model for diverse subset selection. [@kulesza2012dpp] For PSD (K), log-det exhibits diminishing returns and admits greedy approximation guarantees under standard submodularity conditions. [@nemhauser1978submodular] In practice, MMR offered a simpler and more scalable approximation, so it was used in all reported runs.
Stage D — SSDE: Self-Supervised Divergence Embedding (enhanced selector)
The baseline selector uses hand-designed signatures (s_i) both for scoring and for redundancy control. To obtain a more stable low-dimensional geometry for diversity selection, we learn a Self-Supervised Divergence Embedding: [ z_i = \frac{g_\theta(\tilde s_i)}{|g_\theta(\tilde s_i)|2}\in\mathbb{R}^{p}, ] where (g\theta) is a small MLP projection head and (\tilde s_i) is a stochastically perturbed view of (s_i) (feature dropout + Gaussian noise), analogous to “two-view” contrastive learning. [@chen2020simclr]
D.1 Clear definitions (anchor / positives / hard negatives)
For an anchor image (x_i), we define training pairs and negatives as follows (in our implementation, this is done in signature space, for efficiency):
- Anchor: (x_i) (represented by its divergence signature (s_i)).
- Positive: the same image (x_i) under a second stochastic “view” of its signature (e.g., independent feature dropout + noise), yielding (\tilde s_i^{(1)}) and (\tilde s_i^{(2)}), then embeddings (z_i^{(1)}), (z_i^{(2)}). Interpretation: “same stimulus, same cross-model disagreement pattern,” analogous to augmentations in SimCLR. [@chen2020simclr]
- Hard negatives: images (x_j) whose disagreement profiles are confusable with (x_i) in the current embedding geometry (high similarity), but should be separated to prevent SSDE collapse into near-duplicate clusters. We implement this by mining the top-(k) most similar negatives in-batch and upweighting them. [@robinson2021hard_negatives; @kalantidis2020hard_negative_mixing]
D.2 InfoNCE-style contrastive loss (with mined negatives)
| Let (\mathrm{sim}(u,v) = \frac{u^\top v}{ | u | _2 | v | _2}) (cosine similarity) and (\tau>0) be the temperature. For each anchor (i), define a mined negative set (\mathcal{N}_i) (the other examples in the batch), and a hard-negative subset (\mathcal{H}_i\subset\mathcal{N}_i) (top-(k) by similarity). |
A generic weighted InfoNCE loss can be written: [ \mathcal{L}_i =============
-\log
\frac{
\exp(\mathrm{sim}(z_i^{(1)}, z_i^{(2)})/\tau)
}{
\exp(\mathrm{sim}(z_i^{(1)}, z_i^{(2)})/\tau)
+
\sum_{j\in\mathcal{N}i}
w{ij},\exp(\mathrm{sim}(z_i^{(1)}, z_j^{(2)})/\tau)
},
]
where the hard-negative weights are:
[
w_{ij}=
\begin{cases}
1+\alpha_{\mathrm{hard}}, & j\in\mathcal{H}_i,
1, & \text{otherwise.}
\end{cases}
]
In our implementation, this weighting is applied equivalently by adding (\log(1+\alpha_{\mathrm{hard}})) to the logits of hard negatives before the (\log\sum\exp) operation. [@robinson2021hard_negatives]
D.3 SSDE-trained space used for selection
After training SSDE, we perform diversity-aware greedy selection in the learned space. The SSDE-aware marginal score used at selection step (t) is: [ s_{\mathrm{SSDE}}(x_i \mid S_{t-1}) ===================================
d_i
\lambda\cdot \max_{j\in S_{t-1}} \langle z_i, z_j\rangle, ] where (S_{t-1}) is the set selected so far, and (\langle z_i,z_j\rangle) equals cosine similarity because (z) vectors are normalized.
This is exactly the MMR criterion, but the redundancy term is now measured in the SSDE geometry rather than in the raw hand-crafted signature geometry.
Results
Performance scores (higher is better)
We report the divergence score ((1 - \mathrm{avg\ CKA})) for representative submissions using the methods above, as obtained from the hackathon’s leaderboard.
Table 1 summarizes four representative runs: the first two are baseline (with and without MMR), and the last two are SSDE + MMR under two parameter settings (with the best achieving 0.5447). Additional submissions in the same range were repeats/ablations and are omitted for clarity.
Table 1 — Red Team runs and complete hyperparameter settings.
| Run | Method | Selection objective & space | Parameters (all) | Score |
|---|---|---|---|---|
| 1 | Baseline (Top-(K)) | select Top-(K) by (d_i) | Proxy: (d{=}256), (\alpha{=}0.5), (\varepsilon{=}10^{-3}) • Selection: (K{=}1000), (\lambda{=}0), prefilter (P{=}) N/A, quota: none | 0.4782 |
| 2 | Baseline + MMR | (d_i - \lambda\max_{j\in S}\mathrm{sim}(s_i,s_j)) | Proxy: (d{=}256), (\alpha{=}0.5), (\varepsilon{=}10^{-3}) • Selection: (K{=}1000), (\lambda{=}0.20), prefilter (P{=}20000), quota: none | 0.5099 |
| 3 | SSDE + MMR | (d_i - \lambda\max_{j\in S}\langle z_i,z_j\rangle) | Proxy: (d{=}256), (\alpha{=}0.5), (\varepsilon{=}10^{-3}) • SSDE: (p{=}128), hidden (=512), epochs (=8), batch (=2048), lr (=3\cdot10^{-4}), wd (=10^{-4}), (\tau{=}0.10), mask (=0.20), noise (=0.01), hard-(k{=}128), (\alpha_{\mathrm{hard}}{=}2.0), score_temp (=1.0), max_train (=0), amp (=) on, seed (=0) • Selection: (K{=}1000), (\lambda{=}0.25), prefilter (P{=}20000), quota: none | 0.5409 |
| 4 | SSDE + MMR + quota | same as Run 3 | Proxy: (d{=}256), (\alpha{=}0.5), (\varepsilon{=}10^{-3}) • SSDE: same as Run 3 • Selection: (K{=}1000), (\lambda{=}0.35), prefilter (P{=}20000), quota: ObjectNet 700 / ImageNet 300, seed (=0) | 0.5447 |
Structure of disagreement and diversity
- Selected images are systematically harder: Figure 2 shows the selected set’s hardness distribution is shifted toward higher (u_i), with the selected median around (\approx 0.71), consistent with targeting cross-model disagreement rather than random sampling.
- Model suite is heterogeneous: Figure 3 shows substantial variation in average model–model agreement (some clusters of higher agreement, and many low-agreement pairs), motivating multi-directional disagreement terms and diversity-aware selection.
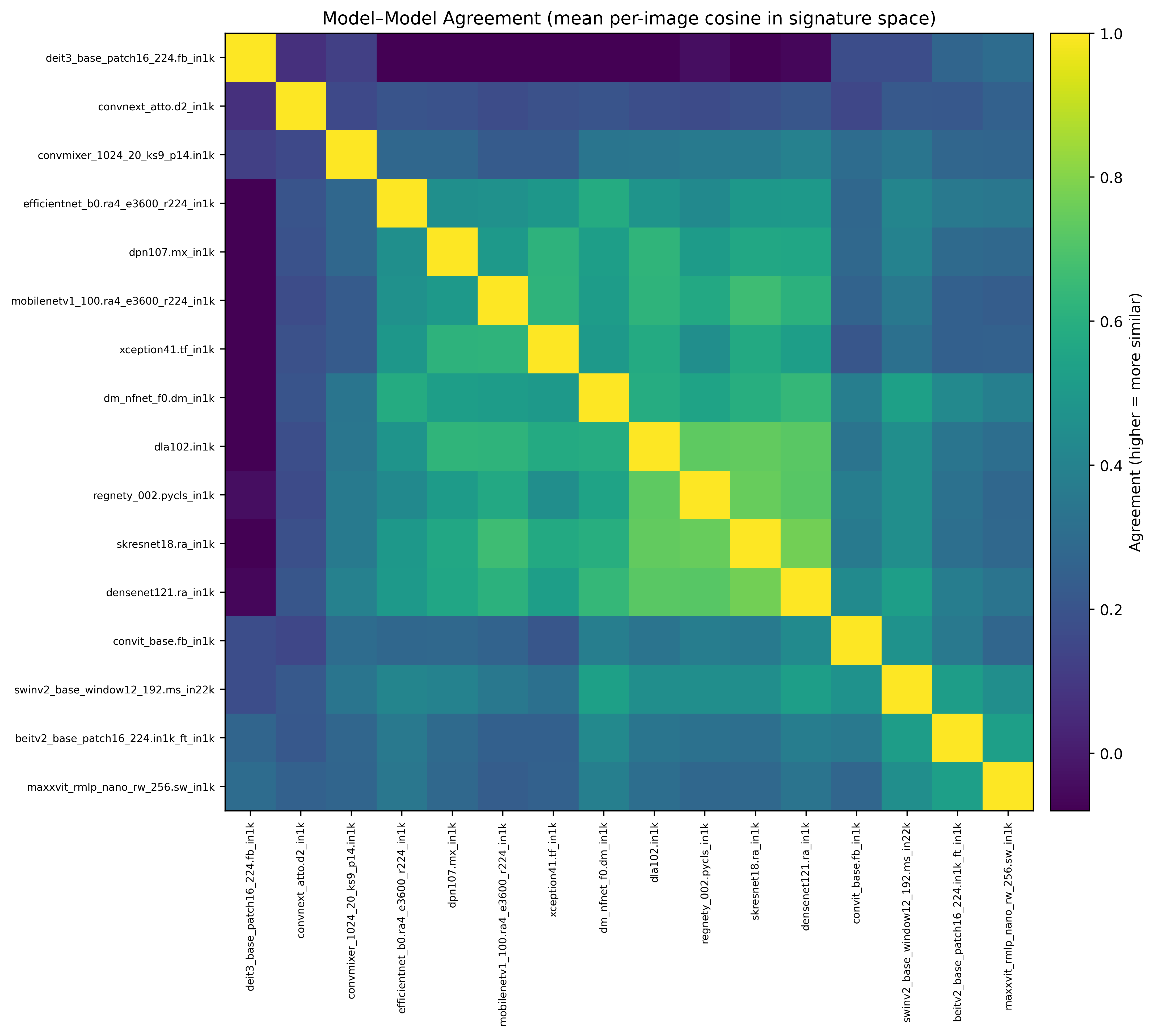
- Selected set remains diverse: Figure 4 visualizes a representative subset RDM (1 − cosine), indicating broad dispersion with limited strong block structure (i.e., reduced redundancy).
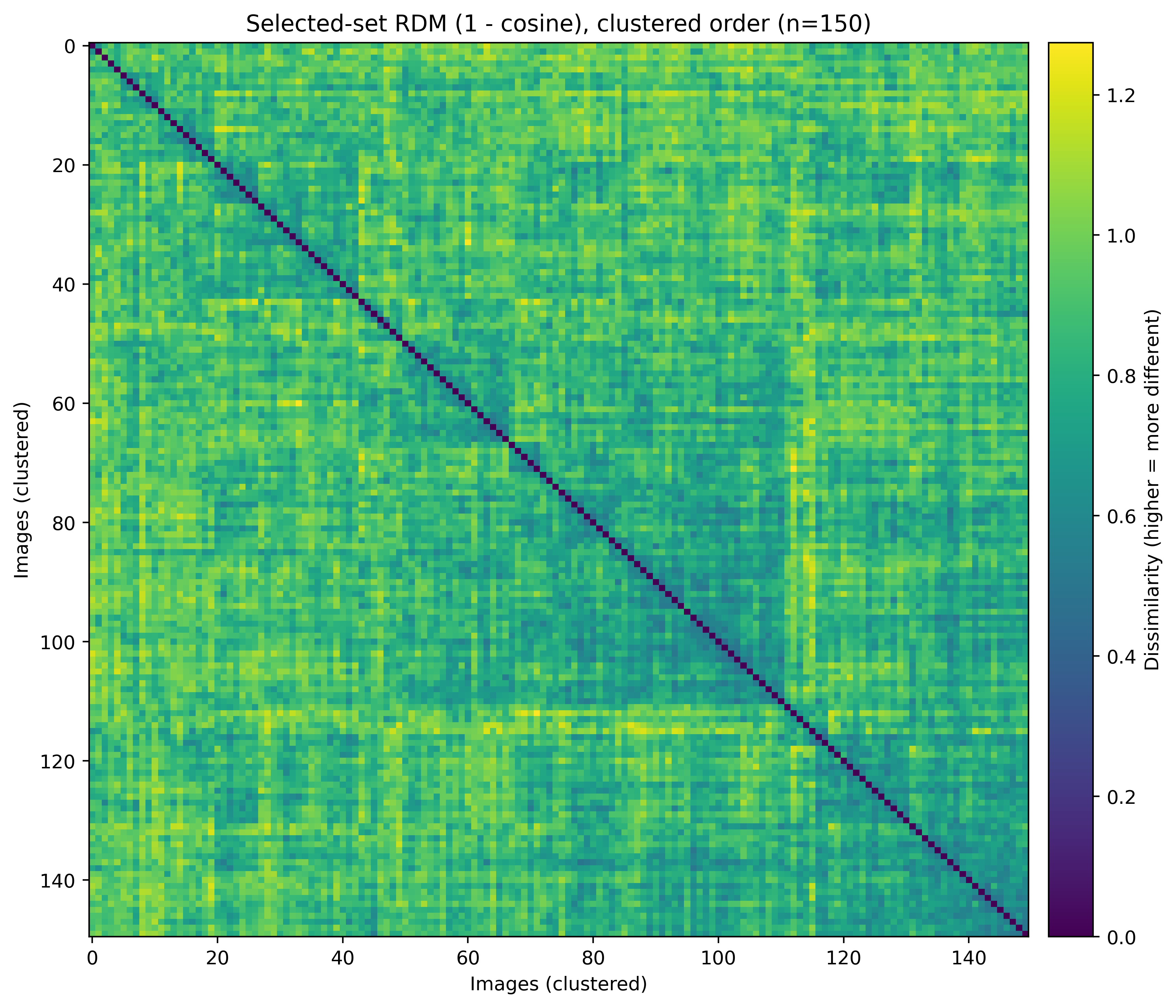
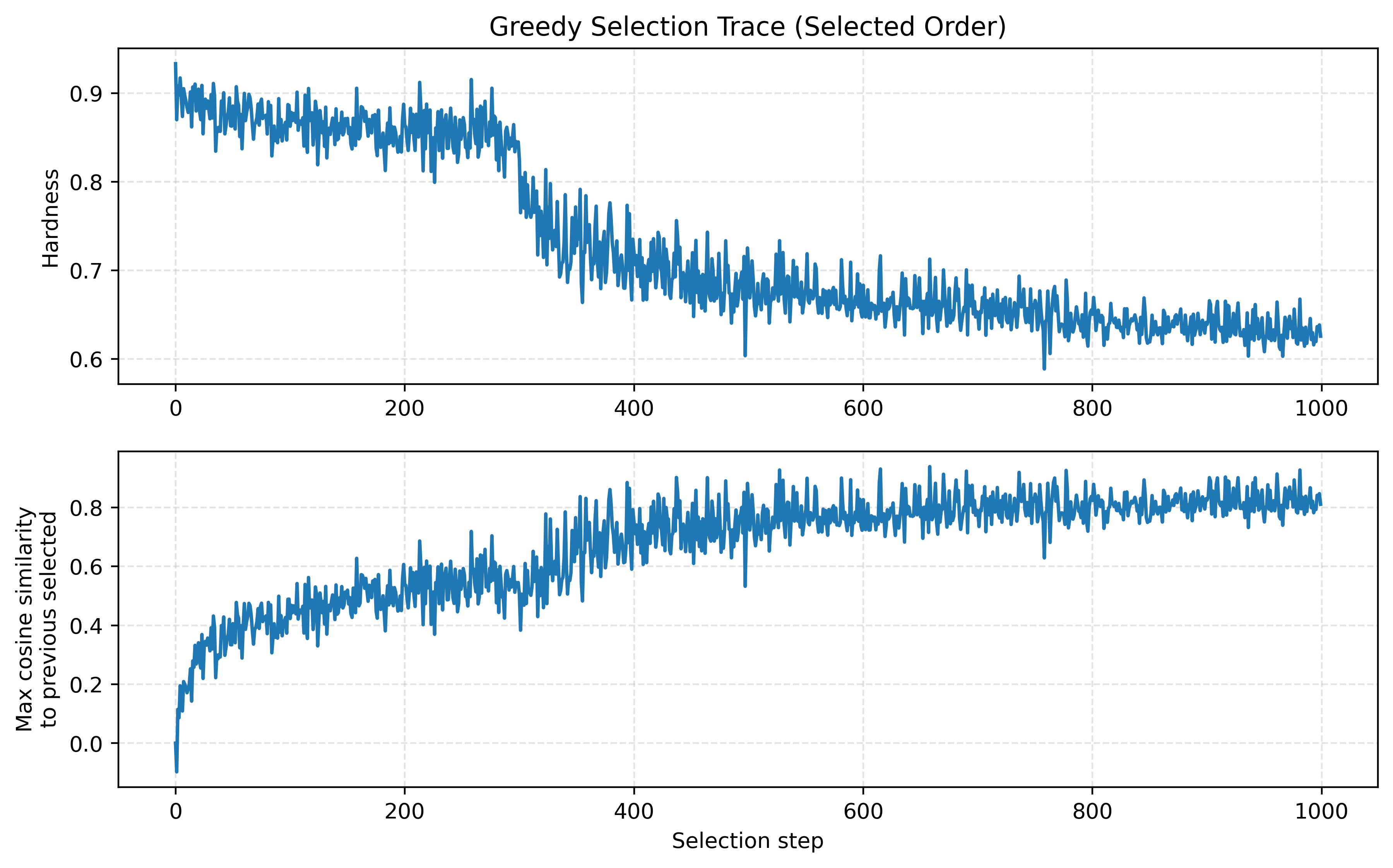
- Greedy trade-off dynamics: Figure 5 shows that as selection proceeds, chosen items tend to become less hard (hardness decreases) while redundancy (max similarity to previous) increases, which is expected for greedy selection balancing signal and coverage.
Observed sensitivity to key experimental knobs
- Diversity weight (\lambda): introducing diversity ((\lambda>0)) improved over Top-(K) (Run 2 vs. Run 1), consistent with the idea that redundant high-score images contribute limited marginal information. [@carbonell1998mmr]
- SSDE geometry vs. raw signatures: SSDE increased scores substantially (Runs 3–4 vs. Run 2), suggesting that a learned embedding geometry for divergence profiles can be a better space for diversity control than hand-engineered signatures alone. [@chen2020simclr; @robinson2021hard_negatives]
- Dataset mixture (quota): the best run used an explicit ObjectNet/ImageNet mixture, consistent with ObjectNet’s goal of probing robustness under dataset bias and distribution shift. [@barbu2019objectnet]
Discussion
Why does SSDE help?
A plausible interpretation is that the pipeline decomposes the divergence objective into signal and coverage:
- The divergence score (d_i) is a signal proxy, designed to prioritize stimuli where models disagree strongly and in multiple directions (via (\ell_i)).
- SSDE provides a learned geometry over divergence profiles that helps distinguish “meaningfully different” disagreement modes from near-duplicates; hard negatives are especially helpful for separating close-but-nonidentical signatures. [@robinson2021hard_negatives]
- MMR then enforces coverage across the learned disagreement modes, which can generalize better to the hidden evaluation split than simply maximizing a proxy on the observed pool.
This emphasis on probing representational geometry rather than only accuracy mirrors core themes in representational alignment research. [@sucholutsky2023getting_aligned; @kriegeskorte2008rsa]
Connection to BrainScore-style model comparisons
BrainScore studies compare models by how well their internal representations predict neural/behavioral measurements and show that architectural and training differences can yield substantial representational differences even when models achieve similar task performance. [@kubilius2019brainscore; @schrimpf2020integrative_benchmarking] This supports the general intuition behind this divergence maximization task: stimuli that expose inductive-bias differences (including distribution shifts such as ObjectNet) are likely to magnify representational divergence across a heterogeneous model suite.
Limitations and future work
This was a hackathon setting with limited time, with the goal of improving scores while understanding how algorithmic and modeling choices affect divergence.
- Proxy mismatch: (d_i) is only a proxy for the hidden CKA-based evaluation, so selection can still overfit to proxy artifacts.
- Lightweight SSDE model: SSDE uses a small MLP on divergence signatures (not raw pixels) to keep training fast; while effective, it may not capture higher-order semantic structure that larger self-supervised encoders can learn from images. Future work could combine SSDE with embeddings from modern image self-supervised models (e.g., DINO-style ViTs) to define richer redundancy measures or to mine hard negatives at the pixel/semantic level. [@caron2021dino]
- Scaling: additional compute would allow (i) larger candidate pools, (ii) longer SSDE training with larger batches (often beneficial in contrastive learning), and (iii) ensembling multiple temperatures/seeds for more stable selection. [@chen2020simclr; @he2020moco]
Conclusion
We presented a Red Team stimulus selection pipeline grounded in established representational-analysis and diversity-selection ideas:
- Multi-model divergence signatures that summarize disagreement geometry across models, motivated by representational similarity analysis and alignment frameworks. [@kriegeskorte2008rsa; @sucholutsky2023getting_aligned]
- A multi-directional disagreement proxy incorporating a log-det term to emphasize disagreement spread across multiple directions. [@krause2008sensor_placement]
- Diversity-aware greedy selection (MMR), with conceptual ties to log-det/DPP diversity. [@carbonell1998mmr; @kulesza2012dpp]
- SSDE, learning a compact divergence embedding via contrastive learning with hard negative mining, improving diversity control in a principled way. [@chen2020simclr; @robinson2021hard_negatives]
Across the reported runs, SSDE + diversity selection produced the best score (0.5447). These results should be interpreted cautiously due to proxy mismatch and limited hyperparameter exploration. With more time and GPU budget—especially to broaden the candidate pool and to explore richer self-supervised image features—the same methodological framework could plausibly yield further improvements.
Appendix
A. CKA / HSIC relationship
Given centered representation matrices (X\in\mathbb{R}^{n\times p}) and (Y\in\mathbb{R}^{n\times q}), linear CKA can be expressed in terms of HSIC: [ \mathrm{CKA}(X,Y)= \frac{\mathrm{HSIC}(X,Y)}{\sqrt{\mathrm{HSIC}(X,X),\mathrm{HSIC}(Y,Y)}}. ] [@kornblith2019cka; @gretton2005hsic]
B. Pseudocode
INPUT:
Candidate images D = {x_i}_{i=1..N} from allowed datasets (ImageNet-val, ObjectNet)
Fixed model suite {f_m}_{m=1..M}
Target set size K = 1000
HYPERPARAMETERS (as used in Table 1):
Projection / proxy:
d = 256 # random projection dimension for model features
alpha = 0.5 # weight for pairwise disagreement u_i vs log-det term ell_i
eps = 1e-3 # log-det stabilization epsilon in log det(G_i + eps I)
Baseline selection (MMR):
lambda_diversity ∈ {0.0, 0.20} # diversity weight
P = 20000 # prefilter top-P by proxy score (for MMR runs)
SSDE training (Run 3–4):
p = 128 # SSDE embedding dimension (proj-dim)
hidden = 512 # MLP hidden dimension
epochs = 8
batch_size = 2048
lr = 3e-4
weight_decay = 1e-4
tau = 0.10 # contrastive temperature
mask_p = 0.20 # feature dropout prob for signature views
noise_std = 0.01 # gaussian noise std for views
hard_k = 128 # top-k hardest negatives per anchor
hard_alpha = 2.0 # hard-negative weight: w = 1 + hard_alpha
score_temp = 1.0 # score weighting temperature (sigmoid) during training
max_train = 0 # 0 means train on all N signatures
amp = on # mixed precision
SSDE selection:
lambda_diversity ∈ {0.25, 0.35}
P = 20000
quota_best = {objectnet:700, imagenet:300} # used only for best run
ALGORITHM:
(1) Representation extraction:
For each image x_i and each model m:
extract layer feature h_m(x_i) ∈ R^{d_m}
(2) Standardize + random project:
Define standardized vector \tilde{h}_m(x_i) (e.g., centering + L2 norm)
e_m(x_i) = P_m * \tilde{h}_m(x_i) ∈ R^{d}
(3) Build per-image matrix:
E_i = [e_1(x_i); ...; e_M(x_i)] ∈ R^{M×d}
(4) Compute disagreement proxies:
u_i = mean pairwise cosine distance across rows of E_i
G_i = (E_i E_i^T)/d
ell_i = log det(G_i + eps I)
score d_i = alpha * u_i + (1-alpha) * clip(ell_i)
Construct signature s_i from flattened upper triangle of G_i + summaries
(5) Train SSDE on signatures (SSDE runs):
Create two signature views per sample with (mask_p, noise_std)
Train MLP g_theta with InfoNCE loss (temperature tau)
Mine hard negatives: top-k by similarity; upweight by (1 + hard_alpha)
Output normalized embeddings z_i = normalize(g_theta(s_i)) ∈ R^p
(6) Select K images (baseline or SSDE):
Prefilter to top P by d_i (if using MMR)
Greedy selection for t = 1..K:
choose i* = argmax_{i not in S} [ d_i - lambda_diversity * max_{j in S} sim(v_i, v_j) ]
where v_i = s_i (baseline MMR) or v_i = z_i (SSDE MMR)
enforce dataset quota if specified (best run)
(7) Output the collection of required images:
{"dataset_name": ..., "image_identifier": ...}